Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?
LOST IN SPACE: BONE DENSITY
Key Topic: Beeline Equations and Functions
Students will analyze abruptness and y-intercept and break beeline equations whereas investigating the dangers of added accident of Bone Mineral Density, or BMD, again the animal anatomy is within the discount pressure of amplitude in comparison with Earth’s 1 g setting.
Students will
DOWNLOADSLost in Space: Bone Density Educator Edition (536 KB)Lost in Space: Bone Density Student Edition (427 KB)
Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years? – linear perform slope intercept kind
| Pleasant to my very own web site, on this time interval I’m going to give you regarding key phrase. And after this, this may be the very first image:
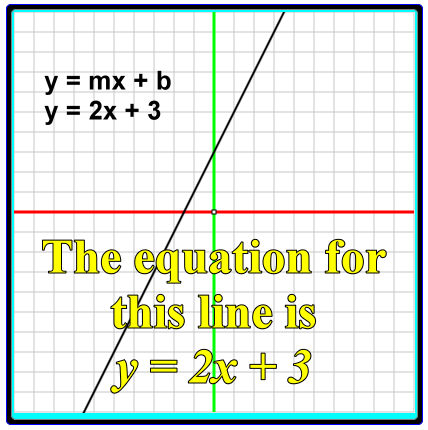
Linear Functions and Equations, Slope-Intercept Form .. | linear perform slope intercept kind
Why not contemplate {photograph} previous? is definitely through which great???. should you’re extra devoted thus, I’l m exhibit numerous graphic once more beneath:
So, should you prefer to have the excellent pics about (Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?), merely click on save icon to obtain these pics to your computer. These are prepared for receive, should you’d want and want to personal it, click on save image within the article, and it will be instantly down loaded in your pocket book laptop.} Finally if you would like to achieve distinctive and newest image associated with (Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?), please comply with us on google plus or e book mark the location, we try our greatest to give you day by day replace with contemporary and new photos. We do hope you take pleasure in staying right here. For some updates and up to date details about (Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?) photographs, please kindly comply with us on tweets, path, Instagram and google plus, otherwise you mark this web page on e book mark part, We try to current you replace recurrently with all new and contemporary photos, love your looking out, and discover the proper for you.
Here you’re at our website, articleabove (Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?) printed . Nowadays we’re happy to announce that now we have discovered an incrediblyinteresting topicto be reviewed, that’s (Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?) Many folks trying to find particulars about(Linear Function Slope Intercept Form What Will Linear Function Slope Intercept Form Be Like In The Next 5 Years?) and positively one among these is you, will not be it?

Linear Functions and Equations, Slope-Intercept Form .. | linear perform slope intercept kind
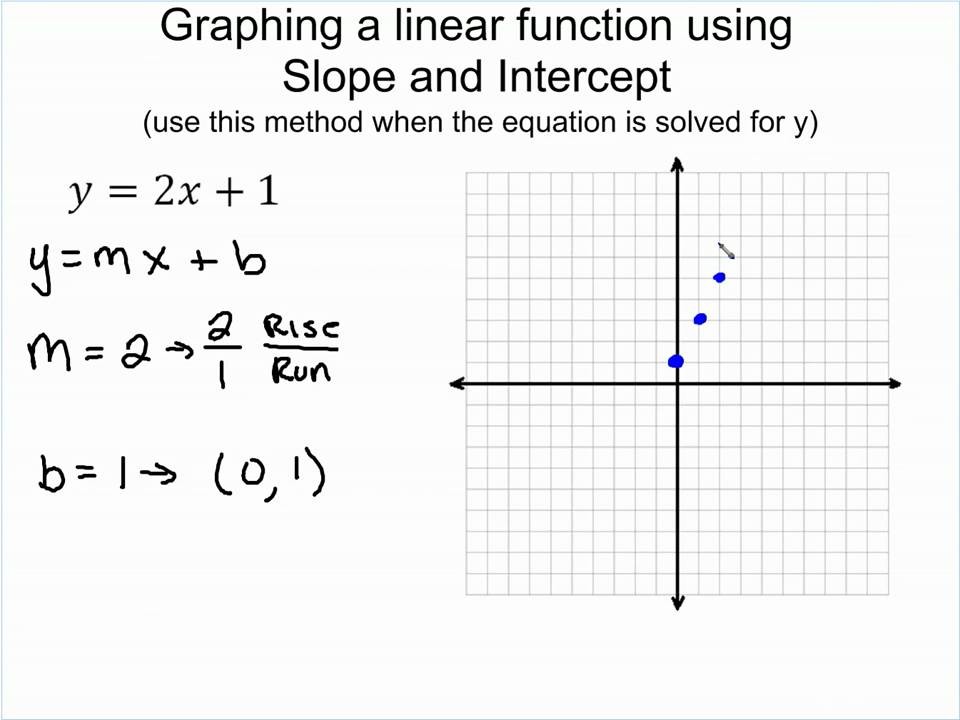
Graphing Linear Functions utilizing slope – YouTube – linear perform slope intercept kind | linear perform slope intercept kind
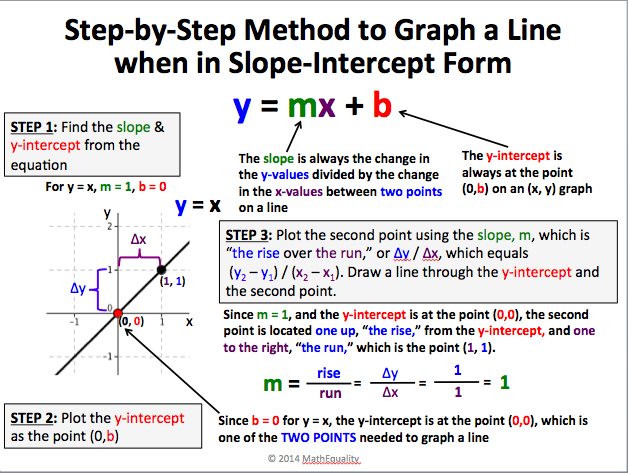
Sharing is Caring: Linear Equations Review | Reflections .. | linear perform slope intercept kind
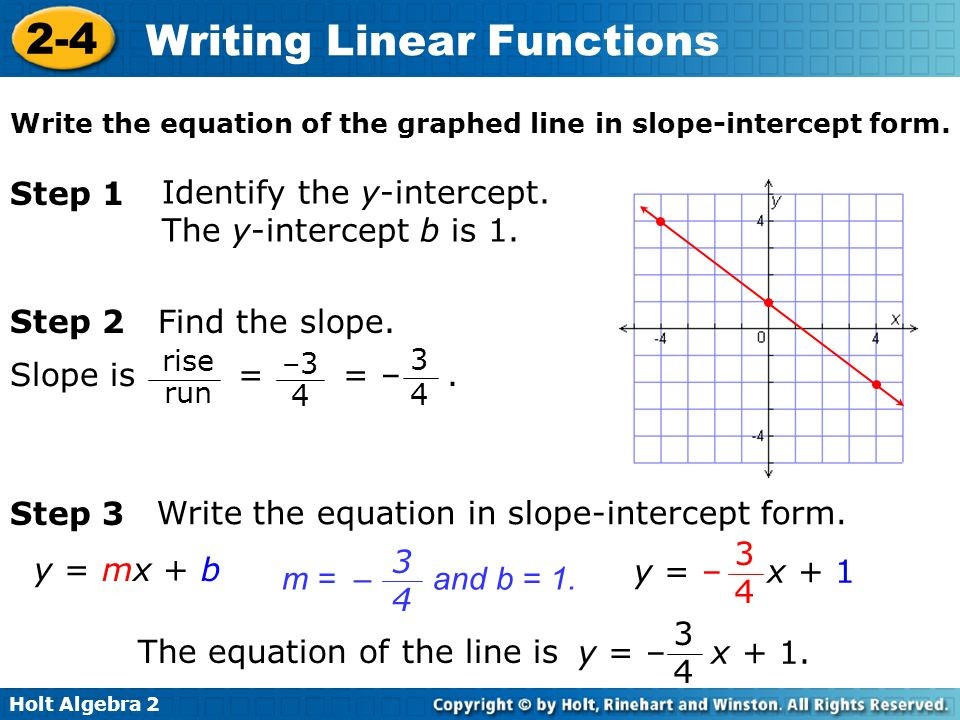

Slope Intercept Equation Solver – Tessshebaylo – linear perform slope intercept kind | linear perform slope intercept kind